 The structure of biological macromolecules is critical to understanding their function, mode of interaction and relationship with their neighbours, and how physiological processes are altered by mutations or changes in the molecular environment.
The structure of biological macromolecules is critical to understanding their function, mode of interaction and relationship with their neighbours, and how physiological processes are altered by mutations or changes in the molecular environment.
Ideally, classical structural biology research should interface more with cellular biology, as it is crucial for the structural data obtained in vitro to be validated within the cellular or tissue context. A true cellular structural biology approach should allow macromolecules to be characterised directly in their native environment. Such an approach would guarantee the high significance of data obtained in vivo or in the cell with the high resolution of a structural technique.
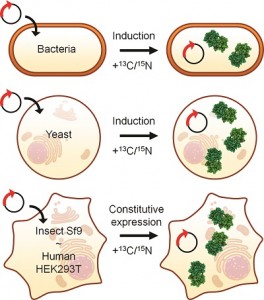
In the Past decade, NMR spectroscopy has been applied to obtain structural and functional information on biological macromolecules inside intact, living cells. The approach, termed “in-cell NMR”, utilises the improved resolution and sensitivity of modern high-field NMR spectrometers and exploits selective enrichment of the molecule(s) of interest with NMR-active isotopes.
Since its inception, in-cell NMR has gradually emerged as a possible link between structural and cellular approaches. Being especially suited to investigate the structure and dynamics of macromolecules at atomic resolution, in-cell NMR can fill a critical gap between in vitro-oriented structural techniques such as NMR spectroscopy, X-ray crystallography and single-particle cryo-EM techniques and ultrahigh-resolution cellular imaging techniques, such as cryo-electron tomography.
In a topical review IUCrJ (2017), 4, 108-118 Lucia Banci and her co-worker Enrico Luchinat, both based at the University of Florence, summarise the major advances of in-cell NMR and report the recent developments in the field, with particular focus on its application for studying proteins in eukaryotic and mammalian cells and on the development of cellular solid-state NMR.